New Study Challenges Assumed Cell Division Process

A new study led by the researchers at Okinawa Institute of Science and Technology Graduate University (OIST) has challenged the assumed aspects of cell division i.e. how this process has been thought to work.
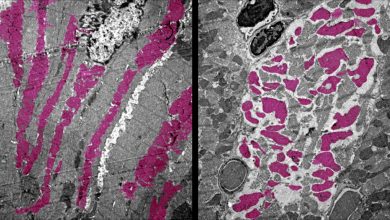
Cell division, essential process of life in which chromosome duplicates into sister chromatid, aligned in an equatorial plane and then segregates into two daughter cells. This process is essential for living things to grow, multiply and make complex structures. But there are still many details which are yet to be explored.
This current study is much focused on cohesion where scientist challenges a key aspect of assumed cell division that how it works.
Cohesin is a protein whose function is to hold the sister chromatid together and letting go when a cell divides and chromosome separate at the last stage of cell division.
Previous studies indicate that the cohesin has ring-like structure encircling the chromatid to hold them together. But when scientist did the study some aspects of this model didn’t add up.
Actually, the cohesin holds the two sister chromatid together and at the time when a cell starts dividing, there is an enzyme called cut1 which opens the ring so that cell can segregate into the daughter cell.
So when the scientist mutated the specific gene CUT1, cell cycle broke down as it couldn’t open the ring-shaped cohesion.
But as normally studied theory suggested if the second mutation is done, it undoes the function of the first mutation and restores cell division, but to our surprise, it didn’t happen. Instead, the cell having both mutations could not break cohesin. With this study in mind, the author concluded that maybe the cohesin did not have ring-like structure because cut1 enzyme needs cohesin’s ring-like structure to function correctly.
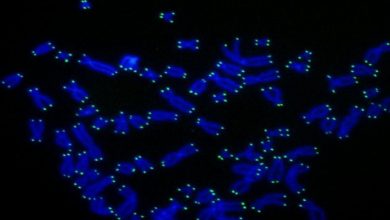
Previous studies in 2003 by Prof. Yanagida’s group had examined some aspects of cohesin’s structure at an atomic level using an extremely high-resolution microscope called an Atomic Force Microscope (AFM), but those results didn’t reveal all the fine details of the complex’s shape. To get a better picture, the OIST team had to look at the genetics.
During their research, scientist sequenced the whole genome of cohesin to find the protein’s part where mutation takes place and examined how it is affecting the structure of the cohesin complex.
They found that the mutation was at head and hinge region of cohesin which joins two main unit of protein. According to previous studies, these regions are located on opposite side of ring structure but genomic analysis reveals that they are located next to each other which make cohesin ring structure questionable.
Therefore researchers proposed that cohesin has a more like a jaw structure to hold chromatid at their place and opens when chromatid needs to move. This model does not need a CUT1 enzyme.
The cohesion is more likely have a tadpole-like structure with one large end and other thinner tapering end.
Dr. Nakazawa and Dr. Xu still don’t have enough evidence to pin down cohesion’s shape. They also still don’t know how cohesin binds to chromatin to hold it in place and whether a single complex or a pair of them is needed to do so. As the team explores these possibilities, they may get closer to proving their new theory – and perhaps even glean a better understanding of diseases caused by abnormal cell division.